Total RNA was extracted according to the manufactures’ protocol and the precipitated RNA was solved in RNAse free water 
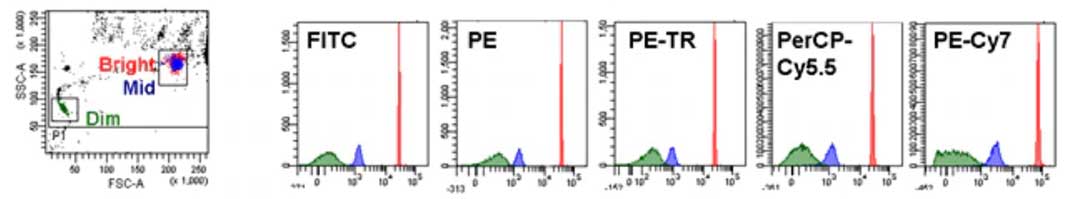
Cells were sorted using a FACSAria III cell sorting system (BD), equipped with FACSDiva software (BD). Dead cells were excluded using 7 amino-actinomycin D (7-AAD) (BD Biosciences). 
Fc receptors were blocked with mouse SeroBlock FcR (CD16/CD32 eBioscience) and cells were stained with FITC-conjugated anti-CD45 (eBioscience), APC-conjugated anti-CD11b (Miltenyi Biotec), and PE conjugated anti-F4/80 (Bio-Rad Laboratories), and allophycocyanin-conjugated anti-CD115 (eBioscience) in PBS/1% BSA/2 mM EDTA. Cells were passed through 70 µm and 40 µm cell strainer, and washed with PBS/1% BSA/2 mM EDTA. No treatment, after sorting cells were directly processed for RNAseqĬell isolation protocol: Single-cell suspensions of wound tissue were prepared by a combination of enzymatic digestion (Liberase Blendzyme, Roche Applied Science) and mechanical disruption (Medimachine System, BD Biosciences) as previously described. GEO help: Mouse over screen elements for information.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
